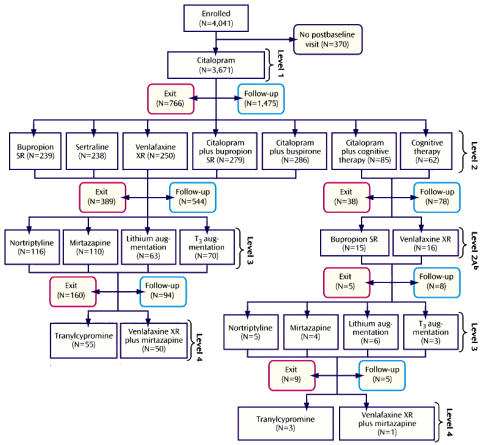
 STAR*D was an elaborate NIMH study aiming to define some sequencing method that would improve the response to antidepressants using the algorithm on the left [as seen from space]. The main outcomes were a database mined for several hundred publications, a self-rating depression scale [QIDS-SR], and a template for future studies called naturalistic at the time – no control group, no blinding, and progress was monitored by a telephone version of the QIDS-SR. From my point of view, it was a thirty-five million dollar misunderstanding…
STAR*D was an elaborate NIMH study aiming to define some sequencing method that would improve the response to antidepressants using the algorithm on the left [as seen from space]. The main outcomes were a database mined for several hundred publications, a self-rating depression scale [QIDS-SR], and a template for future studies called naturalistic at the time – no control group, no blinding, and progress was monitored by a telephone version of the QIDS-SR. From my point of view, it was a thirty-five million dollar misunderstanding…
by Bruce A. Arnow, Christine Blasey, Leanne M. Williams, Donna M. Palmer, William Rekshan, Alan F. Schatzberg, Amit Etkin, Jayashri Kulkarni, James F. Luther, and A. John Rush.American Journal of Psychiatry. 2015 172[8]:743-750.
Objective: The study aims were 1] to describe the proportions of individuals who met criteria for melancholic, atypical, and anxious depressive subtypes, as well as subtype combinations, in a large sample of depressed outpatients, and 2] to compare subtype profiles on remission and change in depressive symptoms after acute treatment with one of three antidepressant medications.Method: Participants 18–65 years of age [N=1,008] who met criteria for major depressive disorder were randomly assigned to 8 weeks of treatment with escitalopram, sertraline, or extended-release venlafaxine. Participants were classified by subtype. Those who met criteria for no subtype or multiple subtypes were classified separately, resulting in eight mutually exclusive groups. A mixed-effects model using the intent-to-treat sample compared the groups’ symptom score trajectories, and logistic regression compared likelihood of remission [defined as a score ≤5 on the 16-item Quick Inventory of Depressive Symptomatology–Self-Report].Results: Thirty-nine percent of participants exhibited a pure-form subtype, 36% met criteria for more than one subtype, and 25% did not meet criteria for any subtype. All subtype groups exhibited a similar significant trajectory of symptom reduction across the trial. Likelihood of remission did not differ significantly between subtype groups, and depression subtype was not a moderator of treatment effect.Conclusions: There was substantial overlap of the three depressive subtypes, and individuals in all subtype groups responded similarly to the three antidepressants. The consistency of these findings with those of the Sequenced Treatment Alternatives to Relieve Depression trial suggests that subtypes may be of minimal value in antidepressant selection.
by Alan F. Schatzberg, Charles DeBattista, Laura C. Lazzeroni, Amit Etkin, Greer M. Murphy, Jr., and Leanne M. WilliamsAmerican Journal of Psychiatry. 2015 172[8]:751-759.
Objective:: The ABCB1 gene encodes P-glycoprotein, which limits brain concentrations of certain antidepressants. ABCB1 variation has been associated with antidepressant efficacy and side effects in small-sample studies. Cognitive impairment in major depressive disorder predicts poor treatment outcome, but ABCB1 genetic effects in patients with cognitive impairment are untested. The authors examined ABCB1 genetic variants as predictors of remission and side effects in a large clinical trial that also incorporated cognitive assessment.Method:: The authors genotyped 10 ABCB1 single-nucleotide polymorphisms [SNPs] in 683 patients with major depressive disorder treated for at least 2 weeks, of whom 576 completed 8 weeks of treatment with escitalopram, sertraline, or extended-release venlafaxine [all substrates for P-glycoprotein] in a large randomized, prospective, pragmatic trial. Antidepressant efficacy was assessed with the 16-item Quick Inventory of Depressive Symptomatology–Self-Rated [QIDS-SR], and side effects with a rating scale for frequency, intensity, and burden of side effects. General and emotional cognition was assessed with a battery of 13 tests.Results:: The functional SNP rs10245483 upstream from ABCB1 had a significant effect on remission and side effect ratings that was differentially related to medication and cognitive status. Common homozygotes responded better and had fewer side effects with escitalopram and sertraline. Minor allele homozygotes responded better and had fewer side effects with venlafaxine, with the better response most apparent for patients with cognitive impairment.Conclusions:: The functional polymorphism rs10245483 differentially affects remission and side effect outcomes depending on the antidepressant. The predictive power of the SNP for response or side effects was not lessened by the presence of cognitive impairment.
To account for the testing of multiple SNPs, SNP p values of 0.05/18<0.0028 ere considered significant using a Bonferroni correction for nine SNPs, each tested for one main effect and one interaction effect. All p values <0.05 are reported for completedness, because of the possibility of false negative results at this significance level.
Remission. In the modified intent-to-treat sample, age [p=0.01] and baseline QIDS-SK score [p<0.001] were significant predictors of remission. Hence, genetic analyses were performed covarying for both.Within the significant overall model [X2=67.58, df=29, p<0.001] only rsl0245483 contributed significantly to prediction of remission. For rsl0245483, there was a significant main effect on remission using multiple testing correction [W=12.64, p<0.001; main effect odds ratio=3.48] and a significant interaction by treatment arm [W=11.18, p=0.001; interaction odds ratio=1.73]. Common allele homozygotes for rsl0245483 responded significantly better to escitalopram [p=0.032] and sertraline [p=0.020] than did minor allele homozygotes. Minor allele homozygotes responded significantly better to venlafaxine [p=0.018]. There were no effects noted in the heterozygotes. The specific contribution of rsl0245483 as a predictor of remission was also verified in univariate models assessing each SNP one at a time.
The effect was similar in whites and nonwhites. In white participants in the modified intent-to-treat sample [N=423], within the significant overall model [X2=61.51, df=29, p<0.001] rsl0245483 had a significant main effect on remission [W=7.22, p=0.007; main effect odds ratio=3.54] and a significant interaction by treatment arm [W=6.99, p=0.008; interaction odds ratio=1.78], which did not pass the multiple testing threshold. For nonwhites, within the significant overall model [X2=55.79, df=28, p=0.001], there was a main effect of rsl0245483 on remission [W=11.42, p=0.001; main effect odds ratio=14.38] and a significant interaction between rsl0245483 and treatment [W=9.81, p=0.002; interaction odds ratio=3.54] that met the multiple testing correction threshold.
My take on this whole line of thinking is suffused with suspicion. It seems to me that the assumption that there will be biomarkers that predict antidepressant response is widely held, but I’m in the dark as to why. At least this study has a hypothesis – the transport of drugs across the blood-brain barrier. This article is accompanied by a Perspective article from the NIMH Intramural Research Program [Clinically Useful Genetic Markers of Antidepressant Response: How Do We Get There From Here?] that suggests [you guessed it] more research.
 TMAP program] and has spawned a steady stream of both public and industry financed research into super-charging antidepressants ever since – now chasing the dream of predictive genetic biomarkers [which might lead to a productive enterprise in commercial testing]. Meanwhile, back at STAR*D, it looks as if someone’s going to have another shot at it:
TMAP program] and has spawned a steady stream of both public and industry financed research into super-charging antidepressants ever since – now chasing the dream of predictive genetic biomarkers [which might lead to a productive enterprise in commercial testing]. Meanwhile, back at STAR*D, it looks as if someone’s going to have another shot at it:Medscape Medical Newsby Kenneth BenderAugust 14, 2015… Somaia Mohamed, MD, PhD, VA Connecticut Health Care System, and coauthors of the article describing the VAST-D study credit the Sequenced Treatment Alternatives to Relive Depression [STAR*D] study for highlighting the frequent inadequate response to initial treatments, but point out that the study did not ultimately identify optimal interventions after initial treatment failure.
They also note that the STAR*D study did not include an atypical antipsychotic augmentation treatment arm, because the study was conducted prior to FDA approval of that indication for an agent in this class.The VAST-D study will incorporate atypical antipsychotic augmentation in the protocol, and the authors indicate that it will answer two principle questions unanswered by STAR*D: "For which patients, under what circumstances, is switching to vs augmenting with other antidepressants the most effective ‘next-step’ strategy, and how does augmentation with atypical antipsychotics compare to either switching or augmenting with antidepressants?"
by Somaia Mohamed, Gary R. Johnson, Julia E. Vertrees, Peter D. Guarino, Kimberly Weingart, Ilanit Tal Young, Jean Yoon, Theresa C. Gleason, Katherine A. Kirkwood, Amy M. Kilbourne, Martha Gerrity, Stephen Marder, Kousick Biswas, Paul Hicks, Lori L. Davis, Peijun Chen, Alexandra Mary Kelada, Grant D. Huang, David D. Lawrence, Mary LeGwin, and Sidney ZisookPsychiatry Research. Published Online: August 05, 2015
Highlights
- Over 2/3s of Major Depressive Disorder cases do not achieve remission on initial treatment.
- Urgent need to identify effective next step treatments for MDD.
- Switching to bupropion-SR vs. augmenting with bupropion-SR or aripiprazole.
- Compare 12-week remission and relapse for up to 6 months after remission.
- Seven methodological issues to balance efficacy and effectiveness.
AbstractBecause two-thirds of patients with Major Depressive Disorder do not achieve remission with their first antidepressant, we designed a trial of three “next-step” strategies: switching to another antidepressant [bupropion-SR] or augmenting the current antidepressant with either another antidepressant [bupropion-SR] or with an atypical antipsychotic [aripiprazole]. The study will compare 12-week remission rates and, among those who have at least a partial response, relapse rates for up to 6 months of additional treatment. We review seven key efficacy/effectiveness design decisions in this mixed “efficacy-effectiveness” trial.
This does seem a poor way to spend $35 million from taxpayers. After reading your 2011 article, I think along the same lines as your thinking early in your training about the nature of depression – i.e. that more often than not it’s difficult life experiences which leads to feelings of depression. Depression is arguably more of a syndrome – i.e. feelings of different degrees caused by different stressors in each individual case – rather than one illness.
In my opinion a study like this is unlikely to work, because it’s trying to compare apples and oranges as if they were all apples, or as if they were three illusory kinds of apples (i.e. melancholic, atypical, anxious). If major depression’s reliability rating is as low as it was in DSM 5 (0.28) and DSM IV (around 0.3 also), what likelihood is there that the subtypes in this study were reliably identified within the participants, if they even exist distinctly in the first place? And what likelihood that such a study would replicate?
I also wonder why they would only follow people up for 8 weeks. Maybe they didn’t have the resources to follow people for a longer period, like say, 2 years. To me it would be much more meaningful to see level of functioning in work and relationships over a 2 year period, rather than symptom controlling for 2 months.
Given such challenges, it’s incredible that these researchers still believe they will identify reliable biomarkers for “depression”. I guess as long as they can get funding to continue this Quixotic quest, they’ll keep trying..
There is still the fundamental problem that just about anything can be called MDD in both practice and research, but waving the bloody shirt of “urgent need” to treat it (meaning its more extreme manifestation) is one sure way to attract funding.
One has to wonder about the diagnoses associated with the genotypes and how “response” was assessed. The use of the Quick Inventory of Depressive Symptomatology–Self-Report (QIDS), which may be administered by telephone, has already been indicted.
The probability is that “response” to an antidepressant will continue to be idiosyncratic, subjective, and highly influenced by environmental factors, despite the added expense of genetic testing to identify a drug match.
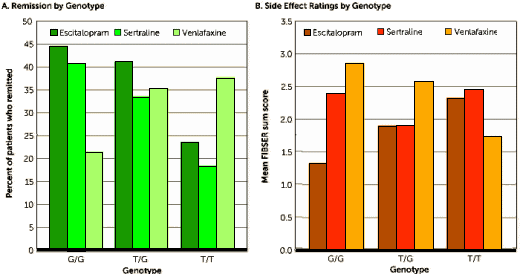
The Schatzberg-Williams report is not encouraging. Much key information is missing. They don’t even tell us how many patients received each of the 3 antidepressant drugs! Plus, they don’t even tell us the distribution of genotypes by drug assignment. Primary remission outcomes are not reported – only percentages of undisclosed denominators (Dr. Mickey’s green Figure A). Variances for the side effect scores are not shown (Dr. Mickey’s brown Figure B). It is also unclear whether the results presented in the Figure and in Table 3 reflect primary observations of outcomes or whether they come from the multivariate, adjusted analyses. The cognitive data for this cohort are never shown, although cognitive impairment is highlighted as an important confound or moderator. These are all lamentable failures of tradecraft. Such is the editorial standard at today’s American Journal of Psychiatry.
Multivariate analyses are reported, supposedly correcting for confounds, i.e., other factors that influenced outcomes. But we are never told how much of the variance is explained by the ABCB1 gene polymorphisms over and above what can be predicted from covariates like site, severity, age, and treatment duration. We are never told the incremental accuracy of predicting remission that results from genotyping.
As for translational science, these authors have no real understanding of why the 3 drugs differed. And one can only look askance at a pharmacogenetic study that fails to report a single blood level of the drugs they are studying. These authors have no clue whether their subjects achieved a pharmacokinetic benchmark (adequate blood levels) that would make all the work of a pharmacogenetic study worthwhile. This iSPOT-D exercise has a gauzy, hand waving quality to it. Does it qualify as solid clinical science? Does it provide a firm basis for “further study… to understand the differences among the agents”? You be the judge.
Thank you Dr. Carroll for that link — although it almost made me lose my lunch! I am SO sick and tired of the Veterans Administration having its precious time and resources siphoned off into research that only benefits private industry, through these so-called Private Public Partnerships. This looks like just one more example.
Check out the Author Affiliations. Alan Schatzberg, affiliated with the Palo Alto VA Hospital! The VA is turning the men and women they serve into guinea pigs. What, they didn’t suffer enough for the sake of corporate interests when they went to Iraq, for Christ’s sake? They have to be force-fed Abilify and Wellbutrin too? (For my money, anyone who runs a study comparing XYZ to Wellbutrin wants to cover up the agitation caused by XYZ by comparing it to something just as bad.)
But what really blew my mind was Schatzberg’s last listed affiliation: Brain Resource, Inc. of San Francisco. What do you make of this? It looks like something even Doctor Oz would be embarrassed to be caught pushing:
http://www.brainresource.com/mybrainsolutions/personal
Johanna,
Look at the second quoted article in this post [ABCB1 Genetic Effects on Antidepressant Outcomes: A Report From the iSPOT-D Trial]. You guessed it. Funded by Brain Resources [“the financing of Australian entrepreneur Evian Gordon’s brain resources”]. Charlie Nemeroff is involved with them too!
It’s the Usual Suspects again. They gravitate towards the big money.